Introduction-Disease pathology and deep learning approach
Acute myeloid leukemia is a severe, mostly fatal hematopoietic malignancy.Transcriptomics
of the NPM gene combined with deep learning could be used as part of an integrated approach
wherein risk prediction, differential diagnosis, and subclassification of AML are
achieved by genomics while diagnosis could be assisted by transcriptomic-based deep
learning.
We find data-driven, high-dimensional approaches—in which multivariate signatures
are learned directly from genome-wide data with no prior knowledge—to be accurate
and robust. Importantly, these approaches are highly scalable with low marginal
cost, essentially matching human expert annotation in a near-automated workflow.
Why deep learning is optimal for this model prediction?
With the increase in the number of datasets,the predition acccuracy will also get increased for deep learning models.Deep learning is a subset of machine learning.
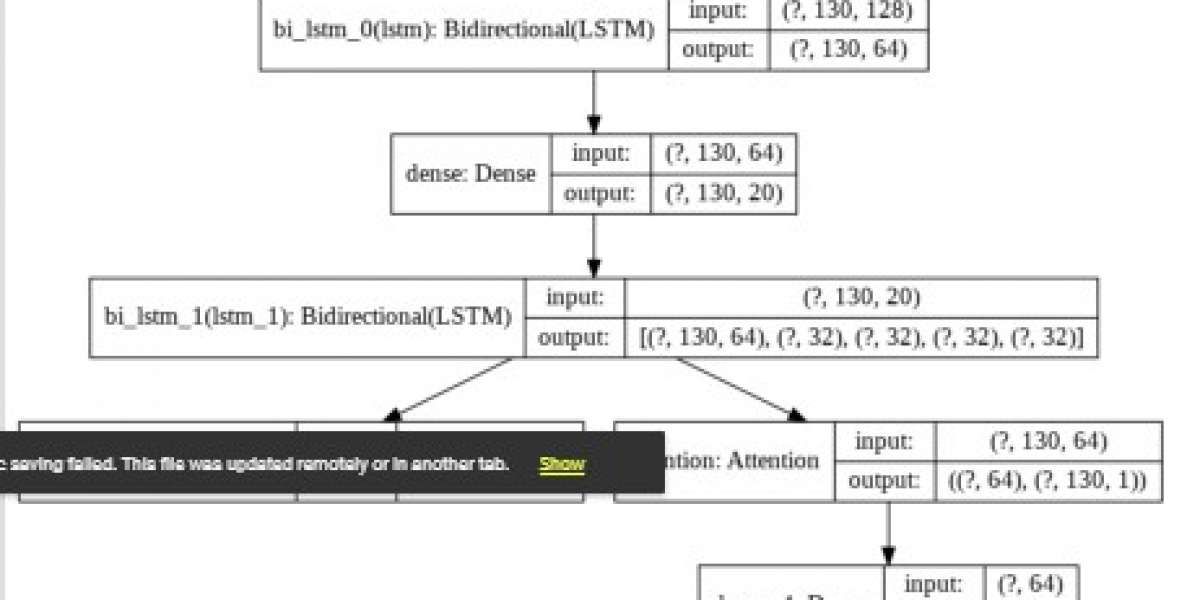
Deep learning technique learn categories incrementally through it’s hidden layer architecture, defining low-level categories like letters first then little higher level categories like words and then higher level categories like sentences.
Each node in the network represents one aspect of the whole and together they provide a full representation of the image. Each node or hidden layer is given a weight that represents the strength of its relationship with the output and as the model develops the weights are adjusted.
Deep Learning algorithms as discussed before are that they try to learn high-level features from data in an incremental manner. This eliminates the need of domain expertise and hard core feature extraction.
Why CNN for this problem?
The number of parameters in a neural network grows rapidly with the increase in the number of layers.The time taken for tuning these parameters is diminished by CNN model.
Dimensionality reduction is achieved using a sliding window with a size less than that of the input matrix for image sets.
The layers of a CNN have multiple convolutional filters working and scanning the complete feature matrix and carry out the dimensionality reduction. This enables CNN to be a very apt and fit network for image classifications and processing.
Overview and uniqueness
AML is characterized by strong transcriptomic signals.The clinical trial findings led
to the suggestion that gene expression profiling could be utilized to define leukemia sub
types and derive useful predictive gene signatures.
- Deep learning based approaches like CNN have the potential for low marginal cost.Based
on the findings and the increasing availability of GEP data derived from peripheral
blood including AML. - We sought to address the subclassification of leukemic disease, where the mutation status
of the leukemic cells is currently the dominant approach as presented in our study
with NPM1 gene’s impact on AML by developing and tuning approaches in which deep
learning tools learn directly from the global transcriptomic data. - After the development of the neural network model,using flask as backend,we can deploy it to the AWS EC2 instances and endpoints.
- Observations from models:
The variations in the NPM proteinis correlated with other attributes like White blood cells count, age ,sex etc.. Majorlydetermine the survival rate of the AML diagnosed with deep learning approach. - For the optimal knowledge of post operations period of survivors, our research and solution
through Neural Networks for not only detecting the effect and amount of NPM1
but also categorizing the type of survivors(disease-free survivors and overall survivors)
proves beneficial.
Handcrafted feature engineering can be eliminated because a deep learning method can
automate this task through the multilayer architecture of a CNN about the rate of remission
of the Cancer cells.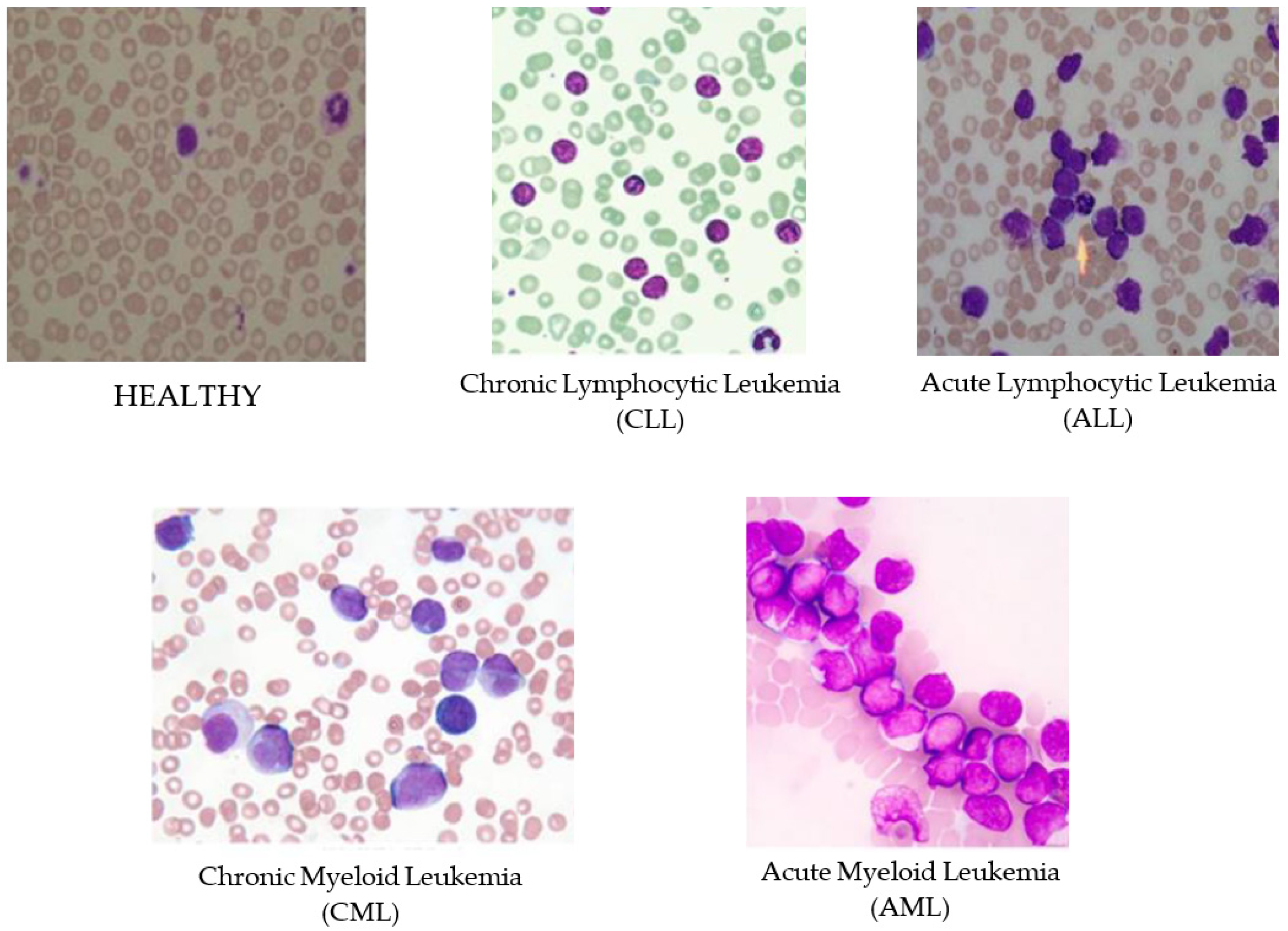
Data sets: Image datasets for AML :
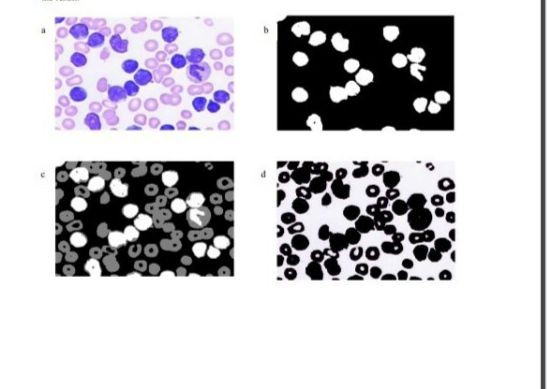
The Acute Lymphoblastic Leukemia Image Database for Image Processing by Fabio Scotti, Associate Professor Dipartimento di Informatica, Università degli Studi di Milano can be used for this project, you can request access by following the instructions on the Download and Term of use page.
The below mentioned dataset can be used for initial modelling for understanding.
https://drive.google.com/file/d/1xn7r9JVCpRZOWEj8gpKA2qFASDhR5izl/view?usp=sharing
Sample data set:
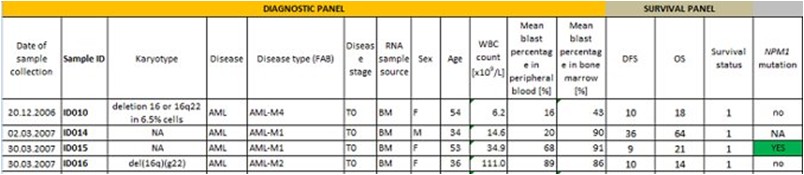
This dataset contains attributes which contribute to the detection of percentage of possibility
of survival after being affected by Cancer(AML – Acute Myeloid Leukaemia)
in people by examining protein NPM transcripts and mutations.
It contains data divided into panels like Mutation panel, Survival panel, RNA-sequence
panel, Diagnostic panel etc..Basically samples are collected from who have been affected
by AML Cancer and have taken treatment.
The variations in the NPM protein is correlated with other attributes like white blood
count, age ,white blood cells count etc.
This dataset provides the necessary diagnosis data for AML cancer prediction with
corresponding survivor rate classification which proves to be in line with the needs of
the proposed problem statement.
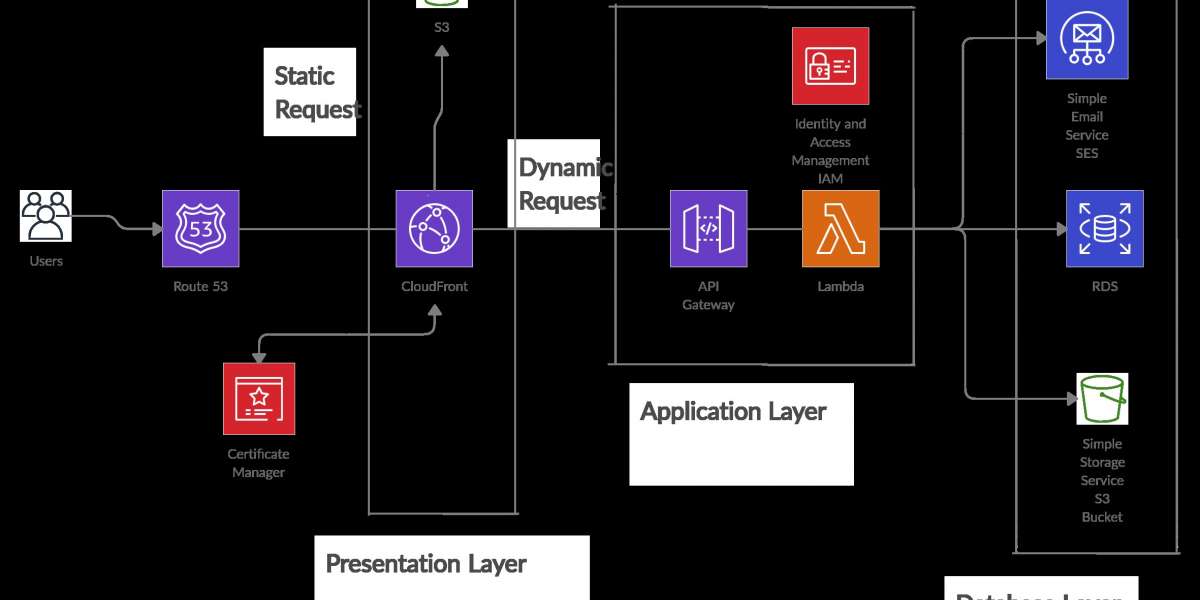
Deploying model in web using AWS EC2 instances and Flask(backend):
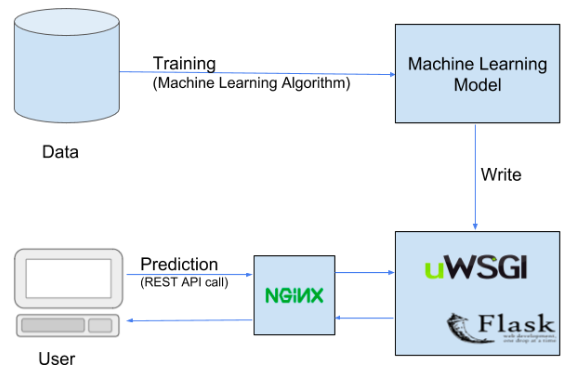
- Train the deep learning model on local system.
- Wrapping the inference logic into a flask application.
- Using docker to containerize the flask application.
- Hosting the docker container on an AWS ec2 instance and consuming the web-service.
Flask is a powerful python microwebserver framework that allows us to build REST API based web-services quickly with minimum configuration hassle.
Containerization
Our should be independent of the underlying machine/OS that runs it. Containerization allows developers to provide such isolation.Containerization of our web-server allows us to avoid the trouble of running into environment related issues. If the containerized code works on one machine, it will surely run on another irrespective of the characteristics of the machine.
Host the Docker container on AWS ECS/EC2 :
Step 1: Set up Amazon ECS.
Step 2: Create a task definition.
Step 3: Configure appropriate service.
Step 4: Configure required cluster.
Step 5: Launch and view generated resources.
Step 6: Open the provided Sample Application.
Step 7: Delete Your Resources.
References:
For Aws hosting:
https://docs.aws.amazon.com/AmazonECS/latest/developerguide/docker-basics.html
[1] Tusher, V. G., R. Tibshirani, et al. (2001). "Significance analysis of microarrays applied to the ionizing radiation response." Proceedings of the National Academy of Sciences 98(9): 5116–5121.
[2] N. Rapin, B. Porse, et al. (2013) Comparing cancer vs normal gene expression profiles identifies new disease entities and common transcriptional programs in AML patients, Blood: 123 (6)
[3] YU Hui, MITRA Ramkrishna, YANG Jing, LI YuanYuan, ZHAO ZhongMing. Algorithms for network-based identification of differential regulators from transcriptome data: a systematic evaluation. SCIENCE CHINA Life Sciences, 2014, 57(11): 1090-1









Nupur Deshpande 4 yrs
Super interesting
Would like to connect and know more!